Multimodal Pooled Genetic Screens Integrating Transcriptomics and Image-Based Phenotypes
Abstract
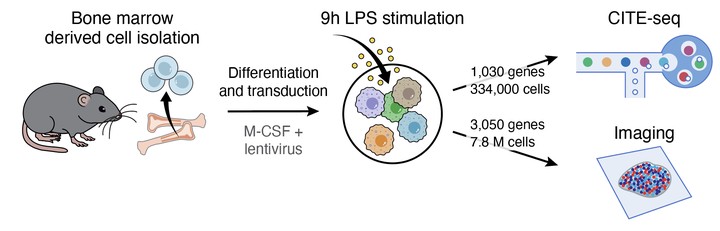
Understanding how genetic perturbations reshape cellular states requires measuring diverse phenotypic modalities at scale. We present PerturbPair, a platform that combines parallel Perturb-Seq and optical pooled screening (OPS) in primary mouse bone marrow-derived macrophages stimulated with lipopolysaccharide—profiling over 334,000 single-cell transcriptomes across ~1,000 gene perturbations and 7.8 million imaging-phenotyped cells across ~3,000 gene perturbations. While RNA and imaging phenotypes were broadly concordant, OPS demonstrated superior sensitivity for weak-effect perturbations and captured post-transcriptional regulatory events that left no transcriptional footprint. To exploit cross-modal relationships, we developed EB-MoCAVI, an empirical Bayesian variational inference framework that imputes RNA profiles for perturbations measured only by imaging and denoises transcriptomic estimates for perturbations with sparse cell coverage. Integrating measured and imputed profiles with rare-variant burden statistics from the UK Biobank identified disease-specific macrophage gene programs and causal regulatory nodes for monocyte counts, type 2 diabetes, and inflammatory bowel disease.